FastQC is a Quality Control Analysis Tool Specifically Designed for High Throughput Sequencing Data
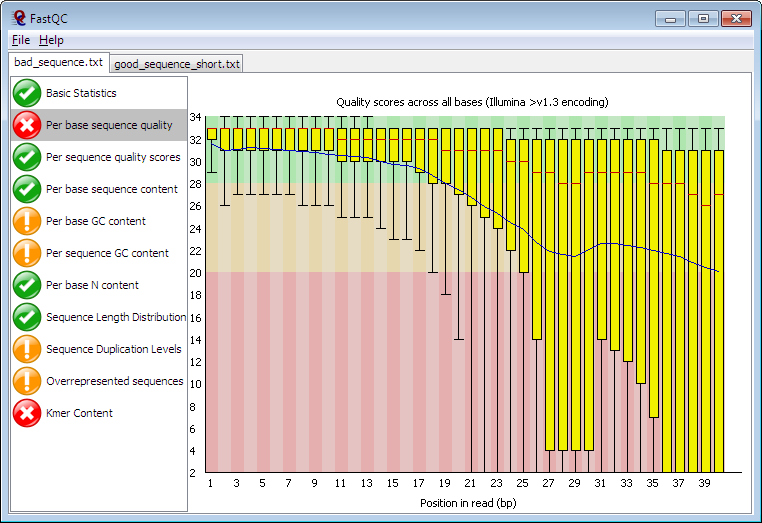
FastQC is a quality control analysis tool for high throughput sequencing datasets. It checks the quality of raw sequence data, runs analyses on sequence files, and produces a report highlighting any unusual areas. It can be used for various types of sequencing techniques.
FastQC will highlight any areas where this library looks unusual and where you should take a closer look.
FastQC is not tied to any specific type of sequencing technique, so it can be used to look at libraries of various experiment types (Genomic Sequencing, ChIP-Seq, RNA-Seq, BS-Seq etc etc).
FastQC is written in Java, making it a cross-platform application that can run on Windows, Linux, macOS, and Unix systems.
Features
- Imports data from BAM, SAM, FastQ, and other compatible file formats
- Provides a comprehensive overview of the data, identifying potential problem areas and suggesting solutions
- Generates detailed summary graphs and tables for a thorough assessment of the data
- Offers the option to export results to HTML, allowing for easy sharing and collaboration
- Supports offline usage, ensuring accessibility and convenience even without an internet connection
Platforms
- Windows
- Linux
- macOS
License
GNU General Public License version 3.0 (GPLv3)