BioImageXD: Open source Microscopy Imaging Software
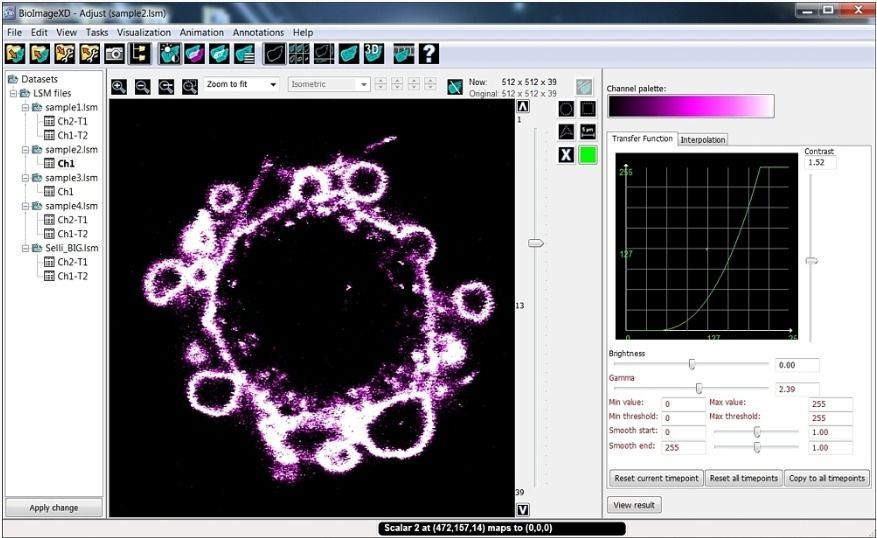
BioImageXD is an open-source microscopy imaging software for processing, analyzing, visualizing, and rendering multi-dimensional microscopy images.
The project was carried out by a team of researchers including microscopists, cell biologists and software engineers from the Universities of Jyväskylä and Turku in Finland, Max Planck Institute CBG in Dresden, Germany and later followed by collaborators worldwide.
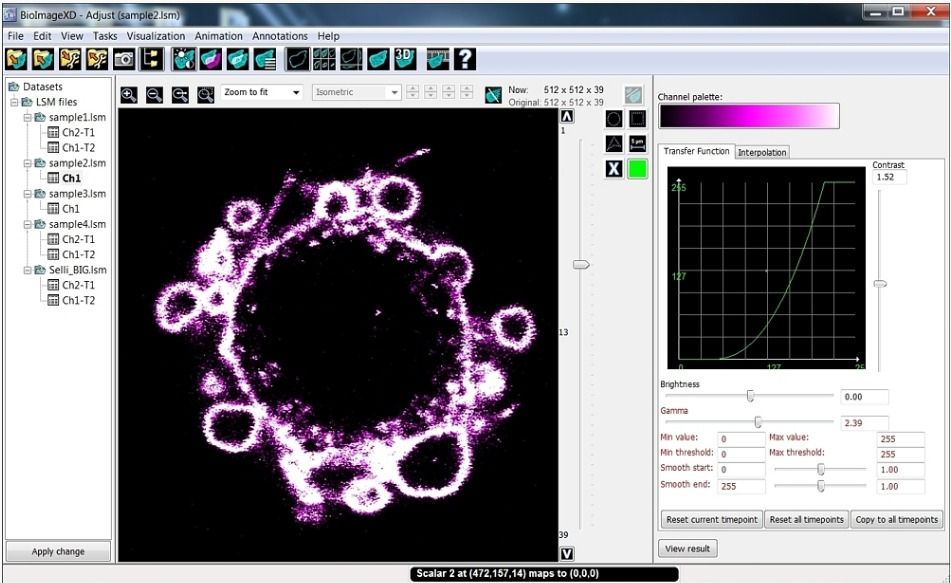
BioImageXD's first version was released in early 2006, the last release was released on July 2012.
The project has positive feedbacks on SourceForge, However, the project has not been updated since 2012.
Highlights
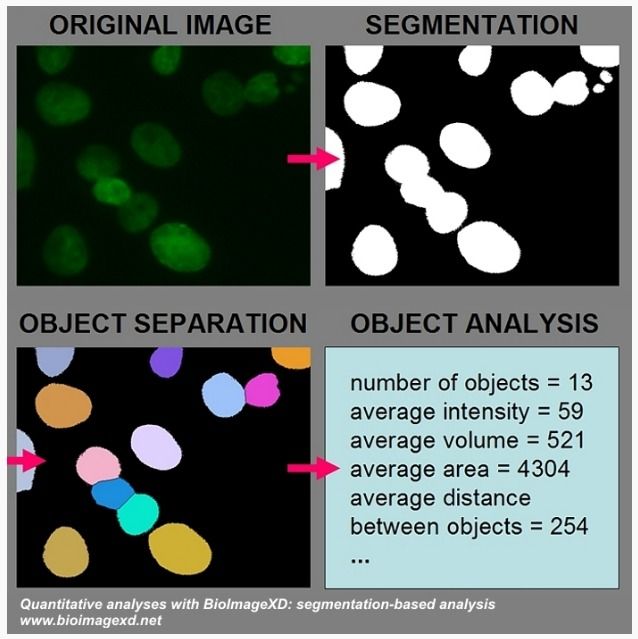

- Free and open source
- All features support 3D time series (4D) data
- Command pipelines and batch processing without programming
- On-screen feature descriptions and color-coding assist
- Leverages VTK and ITK
- Supports many scientific and medical imaging-specific file extensions
- Support scripting with Python
Features
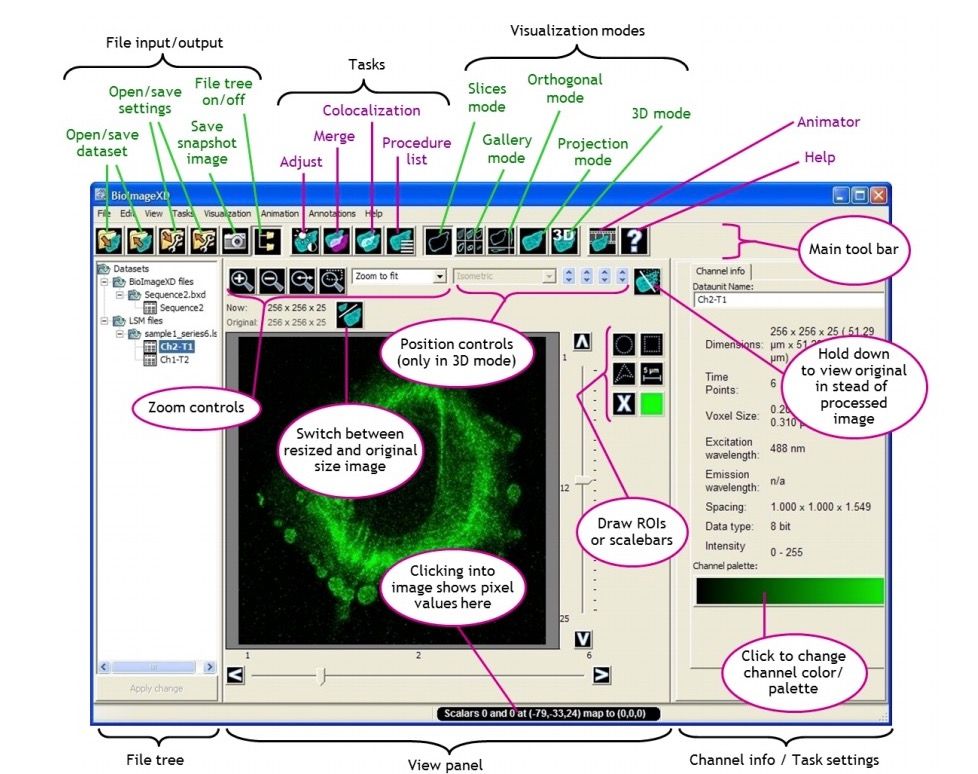
- Multiple Image viewing modes (Slices, Gallery, Orthogonal sections, 3D Renderer, Maximum and average intensity projection)
- Powerful image processing tools C++ powered
- Animator
- 3D surface renderer
- 3D volume renderer
- 3D Ortogonal slices
- 3D protein data visualization
- Axes, angle and distance measurements
- Quantitative analysis: (ROIs analysis, 3D segmentation, Colocalization analysis (voxel and object), Object analysis, Motion tracking, & Fluorescence Recovery After Photobleaching (FRAP) analysis)
- Batch processing support
Supported file types
- Carl Zeiss LSM
- Olympus OIF
- Leica TXT & LIF
- BioRad PIC
- MRC
- Interfile
- TIFF, PNG, JPEG, BMP (read & write)
- OME-TIFF (read & write)
- VTK XML (write)
- BioImageXD internal formats (read & write)
Under the hood
BioImageXD uses C++, Python as main programming languages for the software. It leverages the C++ performance and Python as an interpreted language to run on Windows, Linux, & macOS. It uses several open-source libraries and modules like VTK (The Visualization Toolkit) an open-source BSD-licensed package for 3D computer graphics, image processing, and visualization, & ITK (Insight Segmentation and Registration Toolkit) a cross-platform package for image segmentation and registration.
Prerequisites
- Python 2.7
- wxPython 2.9.3 or newer
- CMake 2.6 or newer
- SWIG 1.3.38 or newer (2.x.x series haven't been tested)
Consideration
- The project has not been updated since 2012
- Does not have official pre-installed packages for the supported platforms (Windows, Linux, & macOS)
- User has to go through installation instruction at the Wiki to install the software
- Requires advanced skills to install, pack, & build.
Conclusion
BioImageXD is a powerful microscopy imaging software, it's unfortunate that it's outdated and not maintained. It also does not have a simple installation procedure or providing pre-configured installation packages for users. However, its layout the features, and specification for open-source developers to learn from it while building similar software.