InVesalius: Open source 3D Medical Imaging reconstruction program
What is InVesalius ?
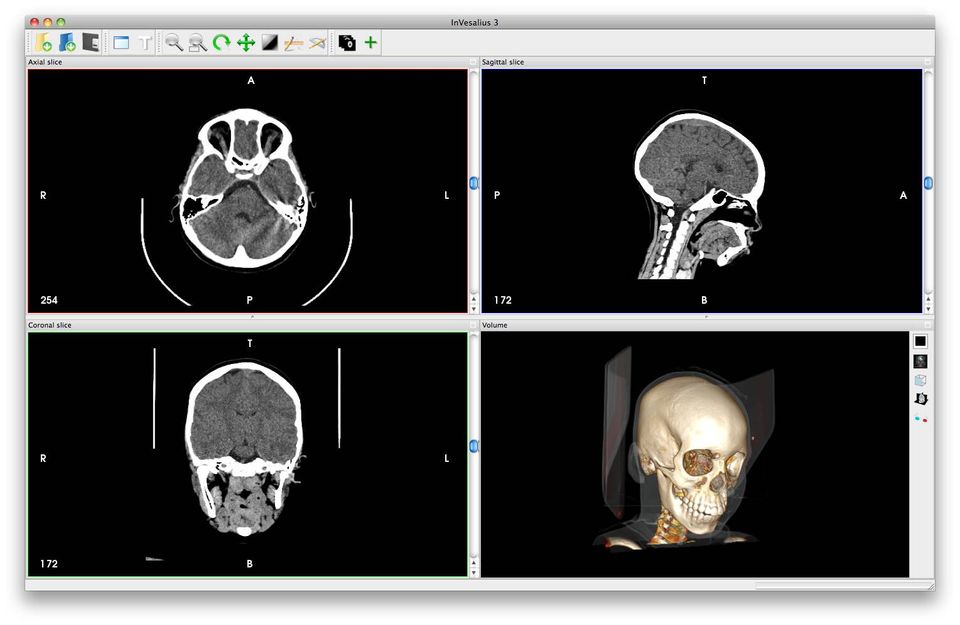
InVesalius Is a free open source 3D medical imaging reconstruction that generates a 3D image from a sequence of 2D DICOM images (CT or MRI). It works for Windows, Linux, & macOS. The project is in active development since 2001, to fulfill the demand for a medical imaging solution for Brazilian hospitals and clinics.
InVesalius is free software, which minimizes the cost for end-users and provides them with advanced features that ease their work. It can be used in medical practice, education, forensic analysis, veterinary, & paleontology.
InVesalius is a production of The Renato Archer Information Technology Center in partnership with the Biomagnetism Laboratory at the University of São Paulo. Founded in 1982 by the ministry of science in Brazil.
The program's interface supports many languages (24 languages) here is some of the languages it supports English, Portuguese, French, Spanish, Turkish, Italian, Czech, Japanese, Catalan, Korean, Romanian and German. It also does provide manuals for end-users in English, & Portuguese.
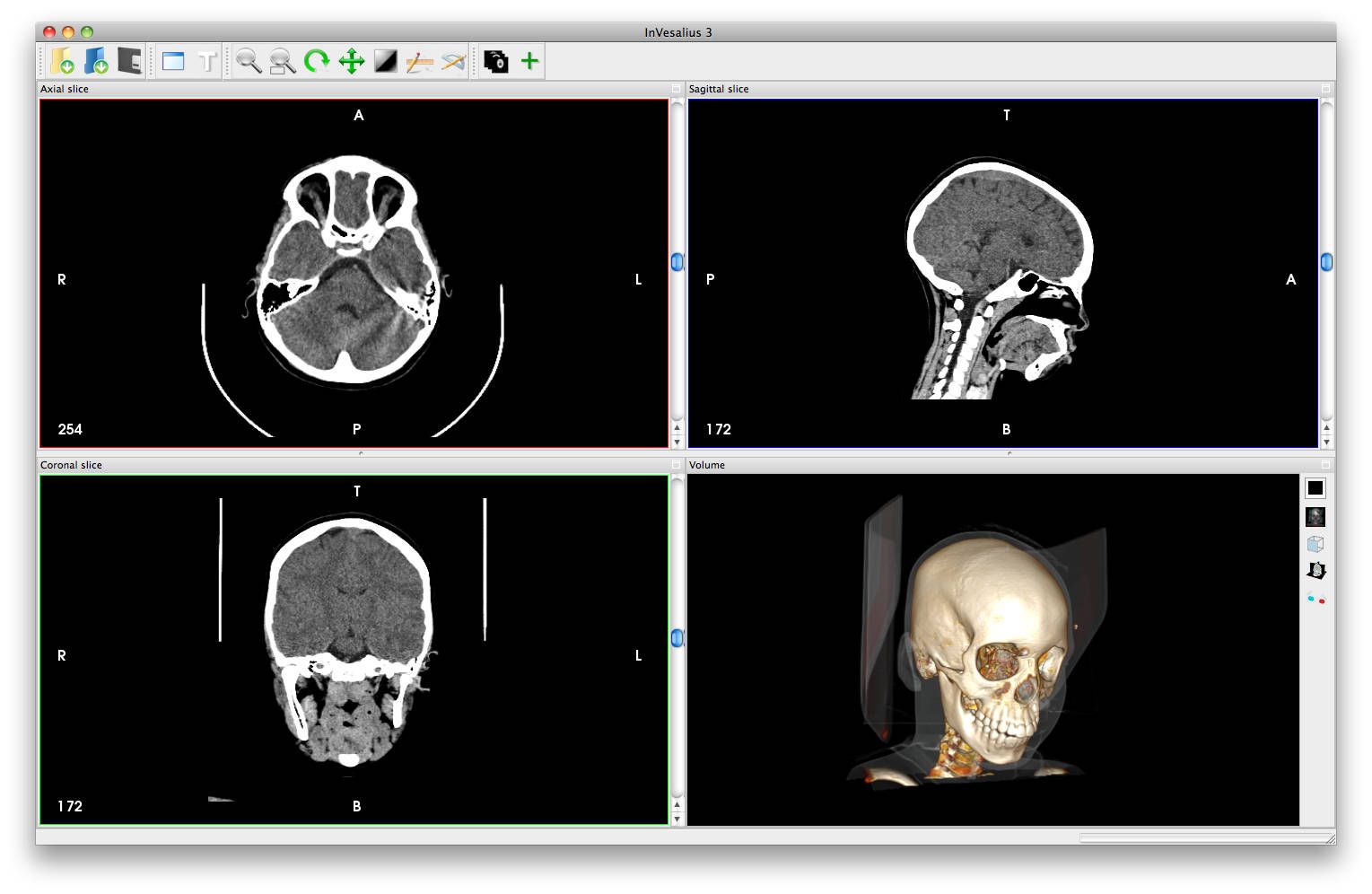
The current new version is 3.1, as it is regularly updated, please keep following the new features, bug fixes, & changed features/ configurations.
InVesalius supports many medical imaging modalities as Computed tomography, magnetic resonance imaging, and microtomography. However, there is a plan to support 3D Ultrasound soon.
Highlights
- Open-source licensed under GPLv2
- Comprehensive end-user manual
- Supports multiple languages
- Easy-to-use interface
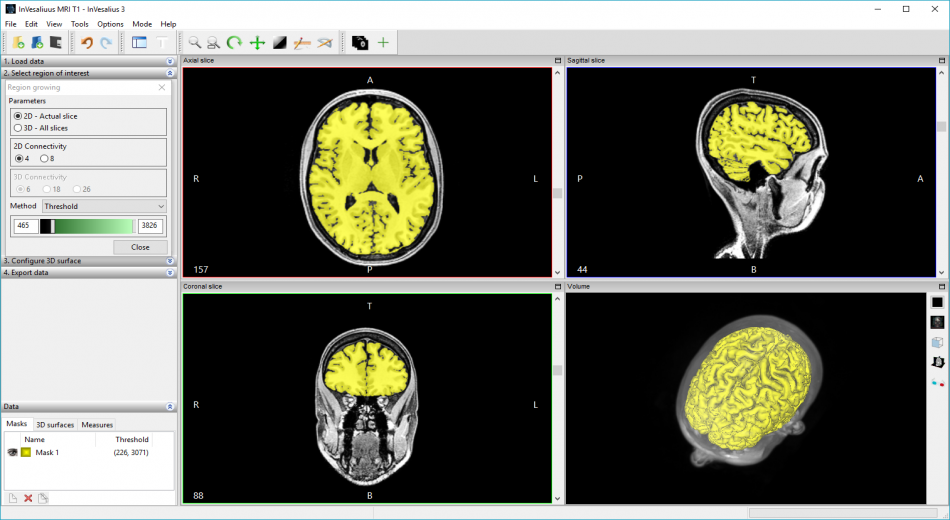
Features
- DICOM support including (a) ACR-NEMA version 1 and 2; (b) DICOM version 3.0 (including various encodings of JPEG -lossless and lossy-, RLE)
- Image manipulation features (zoom, pan, rotation, brightness/contrast, etc)
- Segmentation based on 2D slices
- Pre-defined threshold ranges according to the tissue of interest
- Segmentation based on watershed
- Region growing segmentation
- Edition tools (similar to Paint Brush) based on 2D slices
- Semi-automatic segmentation based on Watershed
- 3D surface creation
- 3D surface connectivity tools
- 3D surface exportation (including binary STL, OBJ, VRML, Inventor)
- High-quality volume rendering projection
- Pre-defined volume rendering presets
- Volume rendering crop plane
- Picture exportation (including BMP, TIFF, JPG, PostScript, POV-Ray)
- Minimum, Maximum or Mean Intensity Projection, Maximum Intensity Difference Accumulation and Contour based visualizations
New Features released in the latest version (3.1)
- Support to open TIFF, BMP, JPEG and PNG files (micro-CT);
- Support to open NiFTI 1 files;
- Support to open PAR/REC files;
- Region Growing based segmentation (Dynamic, Threshold, and Confidence);
- Selection of mask connected components;
- Removal of mask connected components;
- Tool to fill mask holes automatically;
- Tool to fill mask holes interactively;
- Tool to crop mask;
- Support to move slice measure points;
- Menu option to (de)activate the navigation mode;
- Import surface files (STL, PLY, OBJ, and VTP) into InVesalius project;
- Surface area information;
- Segmentation menu with the options: Threshold, Manual, Watershed and Region Growing;
- Created a canvas class (CanvasRendererCTX) to draw 2D forms and text over the vtkRenderer;
- Swap image axes;
- Flip image;
- English documentation.
Supported medical imaging formats
InVesalius supports many medical imaging formats, so as still imaging formats, It supports DICOM, Analyze, PAR / REC, NIfTI, TIFF, JPG, BMP, and PNG formats.
Tutorials
Supported platforms
- Linux: Ubuntu, Debian, Fedora, Arch Linux.
- Windows (32bits, 64bits)
- macOS (10.11 & later) 64 bits
Requirements
Here is a list of the minimum requirements, to run the software:
- Intel Pentium 4 processor or 1.5 GHz equivalent
- 1 GB of RAM
- 80 GB hard drive
- 64 MB graphics card
- 1024 × 768 pixel video resolution
Software install
on Windows
The project supports Windows 32bits/ and 64bits.
on macOS
The project offers macOS a DMG (Mounted Disk Image), which can be installed easily on macOS 10.11 and later
on Linux
There are several packages for (Ubuntu, Fedora, & ArchLinux) and dedicated repo (PPA) for InVesalius for Ubuntu. However, it can be installed in all Linux distro that has Flatpak, With one command in your terminal and you will have the application up and running in on time. Once the install is done the program can run through the system main menu for (Gnome, XFCE, or KDE)
License
The project is released under GNU General Public License v2.0
Contribute
The contribution is not only for developers, but also for anyone to translate user-interface, To join the translation effort, you should signup to Transifex, a cloud-based collaborative translation platform, and contribute in the selected language.
Resources
Developer's note
The end-product depends on many open-source software packages, It uses Python as a core programming language, and several programming libraries that provides image processing, and analysis like Visualization Toolkit VTK, Grassroots DICOM GDCM, Pillow (PIL- Python Image Library Fork)
DICOM/ PACS Viewers Archive
- Open source Free DICOM viewers (Linux, Mac OSX and Windows)
- Free & Open source DICOM Viewers for Mac OSX
- Free DICOM Viewers for doctors: Windows, Linux and Mac OSX
- Open source Browser & Web-based DICOM Viewers
- Free Online Web-based & Cloud DICOM Viewers Services