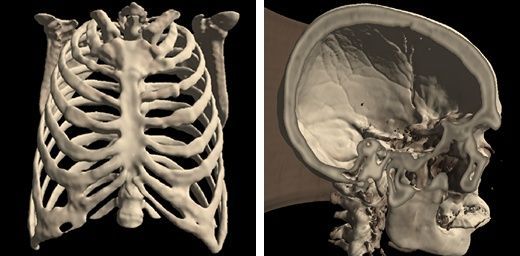
Med3Web: Free, Open source Browser-based DICOM viewer for 2D/ 3D Medical Visualization
Med3Web is an open source free DICOM 2D/ 3D viewer that supports advanced volume and surface rendering & Segmented Data Visualization. It uses web technologies that are compatible and supported by all modern major web browsers on all known operating systems such as HTML5, Javascript, CSS, NodeJS/ NPM. It leverages WebGL feature in browser to render and display advanced 3D models from DICOM Images.
What's WebGL?
WebGL (Web Graphics Library) is a JavaScript API for rendering interactive 3D and 2D graphics within any compatible web browser without the use of plug-ins.
The project is released and maintained by web development agency (Epam.com) among other open-source projects for medical and research usage.
Though Med3Web could be packed as a desktop app, using Electron/ NW projects, however, it was designed and released to be portable, or hosted on the cloud on a web server to be used from browsers. That might be tricky from users with no web developing experience and limited hacking skills, but we will list detailed steps to install it.
Browser support
Currently, only Chrome, Firefox, Safari, Opera browsers are supported. Other browsers can be used with some performance slowdown and other rendering artifacts. Mobile browsers are not currently supported, but some of them can be used with performance/rendering problems and so on.
Requirements
- Modern system
- Modern browsers (Google Chrome, Chromium, Firefox, Opera, Safari)
- WebGL support
- Basic knowledge of coding (NodeJS, NPM) to install
Features
- Open source & Free
- Read data from the local machine, or from a remote server
- Built with HTML5 compatible with all modern browsers
- Works on Modern browsers with WebGL support
- Supports Android Chrome with enabled WebGL
- DICOM, NIfTI, KTX, and HDR formats support.
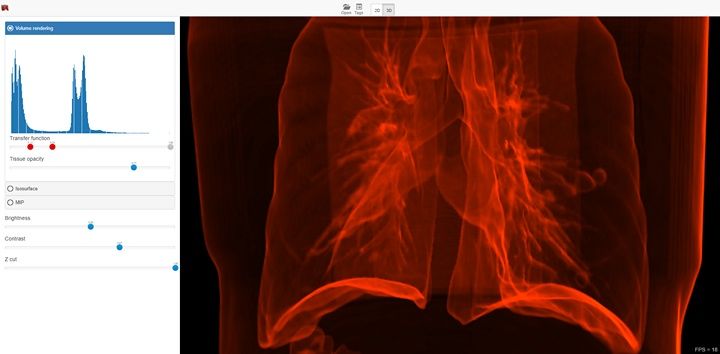
- Real-time adjustment of isosurface and transfer function levels in 3D views.
- Measurement of distances, angles, and areas in 2D views.
- DICOM tag browser
- Real-time volume clipping to open areas of interest
- Automatic volume segmentation
- 5 Mode view support
- Semi-automatic methods for areas of interest opening/selection.
- Segmentation data visualization.
- Advanced 3D visualization (Isosurface Visualization, Volume Visualization, Max Intensity Projection (MIP), Segmented Data Visualization)
- Opens local data and remote data
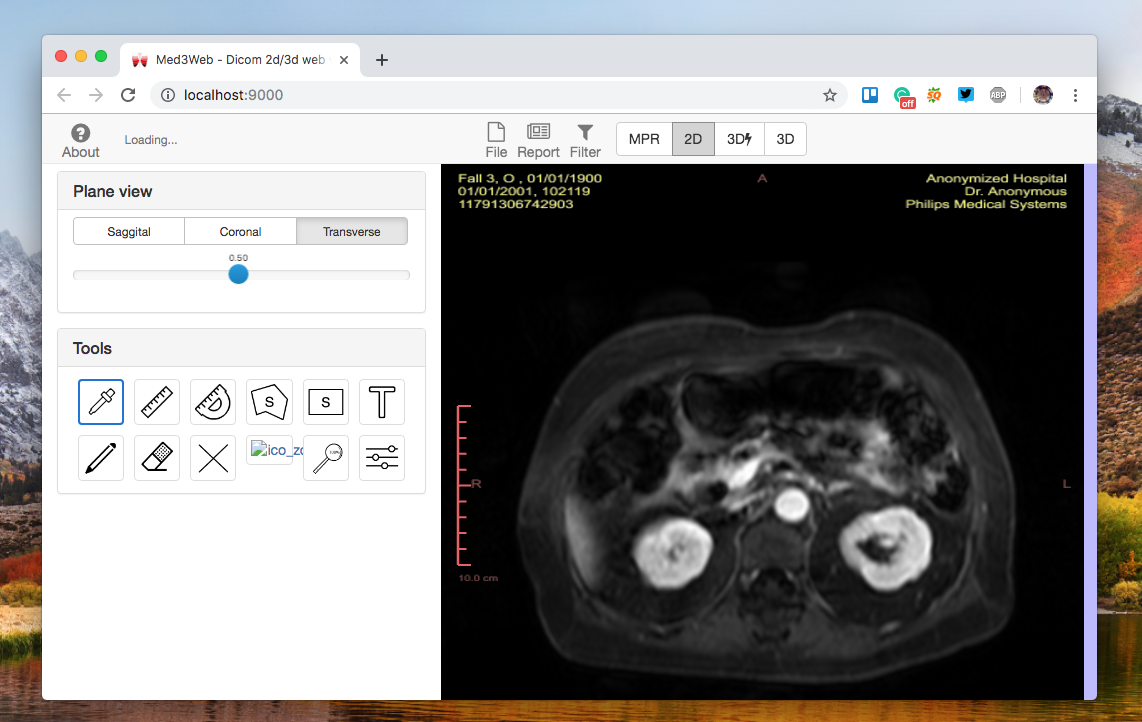
- 2D DICOM viewer
- 3D DICOM viewer
- 3D Volume rendering
- Turbo 3D mode
- Easy setup
- Works without a Server
- Works smoothly on legacy web servers
Technical notes
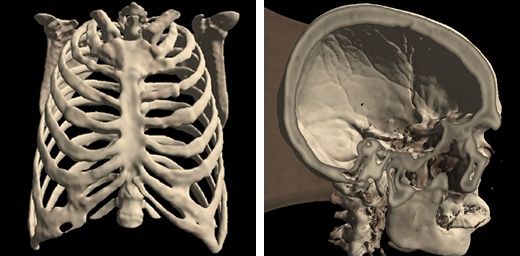
Med3Web is built with pure web technologies, (HTML5, JavaScript, CSS) with WebGL support, using NodeJS for development and NPM (Node Package Manager) for development and building production-ready project. The developers use a set of libraries and framework, DICOM library and visualization tools including
- dicomParser: DICOM parser by cornerstone library, It parses DICOM Part 10 data
- Daikon: A JavaScript DICOM reader.
- Xtk: a Scientific and a medical visualization Toolkit/ framework that utilizes WebGL for advanced 3D medical visualization.
- Three.js: A JavaScript framework for WebGL visualization
Prerequisites
- NodeJS > 6.10.3
- NPM
- gulp
Setup & Install
Open your desktop Terminal
Check for installed NodeJS version, If not installed, please install it
node -v
Clone the project repo
git clone https://github.com/epam/med3web.git
cd med3web
npm install
install gulp
npm install -g gulp-cli
Running the app
npm run start
The app will run quickly and start smoothly even on old machines on those ports
Local: http://localhost:9000
External: http://192.168.1.103:9000
Building for production
Building a static web version, That can be used from the local machine (local server), or installed on a web server.
npm run build
It will build the application in dist directory, however, the static files can't work easily on the local machine, you will need a local server, I would recommend using zero-footprint quick run-up server like http-server,
npm install -g http-server
cd dist
http-server
The app will be ready at port: 8080 "http://127.0.0.1:8080/"
I have tested that on different machines and operating systems including Linux (Solus Linux) with a quite powerful machine with WebGL support, Mac OSX (2011 build with poor WebGL), and on Windows on fairly new gaming PC, It performs very well on modern machines, but poorly on old ones.
License
Licensed under the Apache License, Version 2.0.
Resources
- Med3Web
- Github's page
- Demo app
- WebGL
- Xtk: The X Toolkit: WebGL™ for Scientific Visualization
- ThreeJS: WebGL powered 3D browser-based visualization framework for
- ePam.com The company behind this project
We have covered several DICOM viewers categories:
- Open source Free DICOM viewers (Linux, Mac OSX and Windows)
- Free & Open source DICOM Viewers for Mac OSX
- Free DICOM Viewers for doctors: Windows, Linux and Mac OSX
- Open source Browser & Web-based DICOM Viewers
- Free Online Web-based & Cloud DICOM Viewers Services